ROCKVILLE, MD – Improving the way scientists can see the microscopic structures of the brain can improve our understanding of a host of brain diseases, like Alzheimer’s or multiple sclerosis. Studying these diseases is challenging and has been limited by accuracy of available models.
To see the smallest parts of cells, scientists often use a technique called electron microscopy. Electron microscopy historically involves adding chemicals and physically cutting the tissue. However, this approach can change the way the cells and structures look, perturbing their natural state, and can limit resolution. An alternative method, called cryo-electron tomography (cryo-ET), provides clearer images of the brain's smallest parts in a more native state, however, it requires freezing. Freezing samples to cryogenic temperatures must be done carefully or ice crystals can form, disrupting the native anatomy.
But new research by Benjamin Creekmore in Yi-Wei Chang and Edward Lee’s labs at the University of Pennsylvania shows a new technique to study the human brain's ultrastructure. They will present their research at the 68th Biophysical Society Annual Meeting, to be held February 10 - 14, 2024 in Philadelphia, Pennsylvania.
Creekmore and colleagues obtained brain tissue from autopsies, flash froze it directly on special grids with liquid ethane, and used a powerful tool called a xenon plasma focused ion beam (FIB) to cut thin slices for imaging. This method allowed them to look at the brain tissue in its near-natural state without cutting with a knife blade, adding chemicals, or slower freezing, all of which can lead to changes in the structures.
“The most common way to preserve tissue at a time of autopsy is to put it in a freezer and then use it later. But letting it freeze slowly and then warming it up and then refreezing it also disturbs the tissue. Membranes break and you can lose the normal architecture,” Creekmore explained.
One surprising part of the new method is that it lets them more easily and rapidly freeze much thicker samples—in the past samples were limited to 10 microns using similar approaches. “We were able to freeze samples up to 250 microns thick without ice crystals,” Creekmore said. The process of getting thick samples ready for high-resolution imaging is much faster than with other techniques. This speed-up can allow analysis of a wider array of samples.
By applying this approach to brain tissue from individuals with Alzheimer's disease, they were able to observe intact structures within cells, such as tau fibrils, a hallmark of Alzheimer’s disease, and cellular components that try to break down these fibrils. The team also visualized and measured myelin, a sheath that’s critical for nerve function but that breaks down in certain diseases, such as multiple sclerosis.
“Techniques for imaging human tissue historically have been relatively low resolution and they disturb the native architecture. We wanted to try to come up with a way that we could model and study brain diseases in their natural context,” Creekmore said. Their innovative method provides a first glimpse into the natural state of human brain tissue, offering valuable insights into its anatomy at a high level of detail. This new method can start to provide unique information to determine disease-causing mechanisms of a broad array of brain-related diseases.
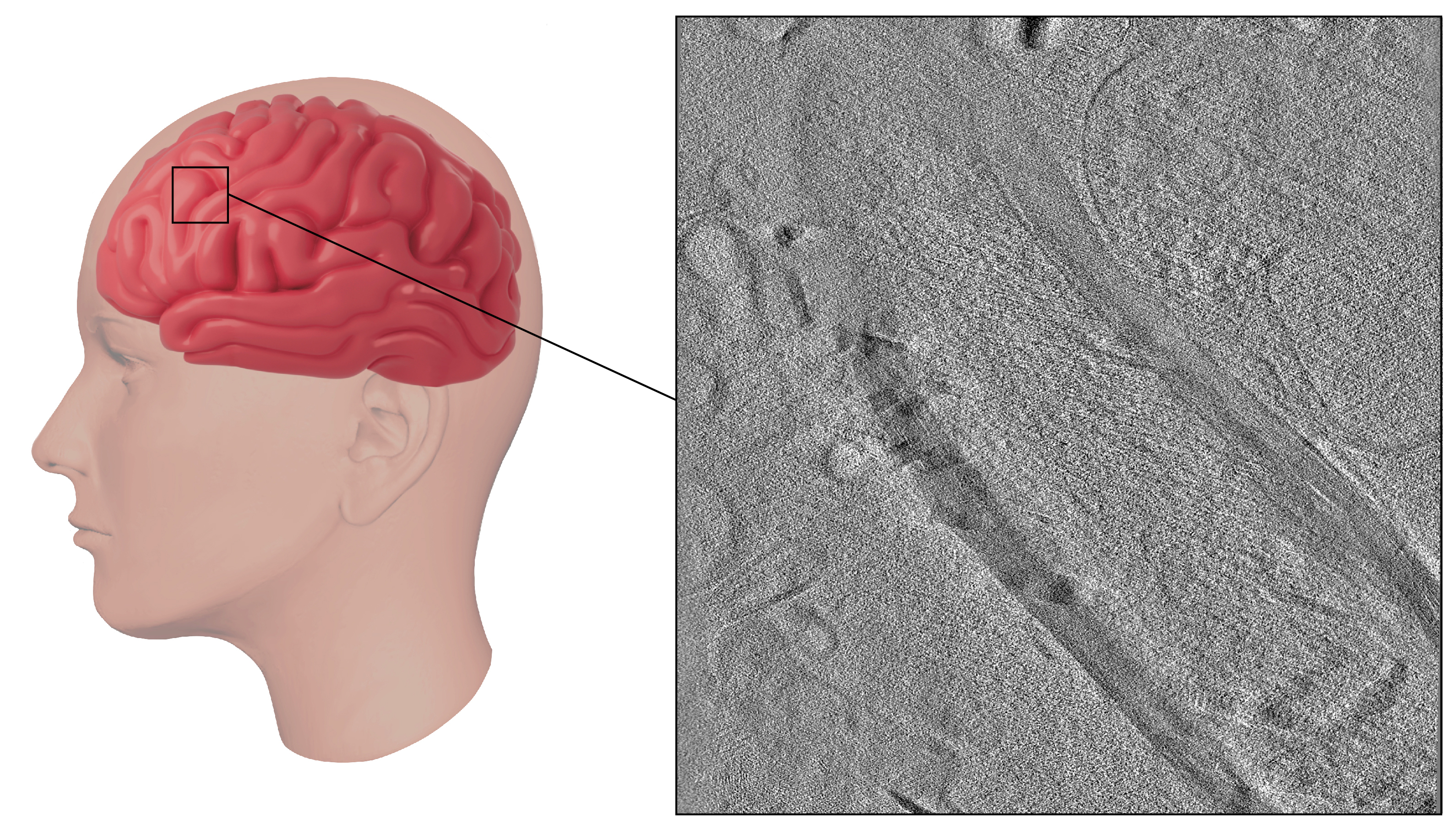
A close-up look at human brain tissue using a new technique developed by Benjamin Creekmore and colleagues.